
Citation: Wei, J.; Hui, A. Review of
Ribosome Interactions with
SARS-CoV-2 and COVID-19 mRNA
Vaccine. Life 2022, 12, 57. https://
doi.org/10.3390/life12010057
Academic Editor: Yih-Horng Shiao
Received: 21 November 2021
Accepted: 28 December 2021
Published: 1 January 2022
Publisher’s Note: MDPI stays neutral
with regard to jurisdictional claims in
published maps and institutional affil-
iations.
Copyright: © 2022 by the authors.
Licensee MDPI, Basel, Switzerland.
This article is an open access article
distributed under the terms and
conditions of the Creative Commons
Attribution (CC BY) license (https://
creativecommons.org/licenses/by/
4.0/).
life
Review
Review of Ribosome Interactions with SARS-CoV-2 and
COVID-19 mRNA Vaccine
Jiao Wei and Aimin Hui *
Shanghai Fosun Pharmaceutical Industrial Development, Co., Ltd., 1289 Yishan Road, Shanghai 200233, China;
weijiao@fosunpharma.com
* Correspondence: aimin.hui@fosunpharma.com
Abstract:
Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) is the causing pathogen
of the unprecedented global Coronavirus Disease 19 (COVID-19) pandemic. Upon infection, the virus
manipulates host cellular machinery and ribosomes to synthesize its own proteins for successful
replication and to facilitate further infection. SARS-CoV-2 executes a multi-faceted hijacking of the
host mRNA translation and cellular protein synthesis. Viral nonstructural proteins (NSPs) interact
with a range of different ribosomal states and interfere with mRNA translation. Concurrent mutations
on NSPs and spike proteins contribute to the epidemiological success of variants of concern (VOCs).
The interactions between ribosomes and SARS-CoV-2 represent attractive targets for the development
of antiviral therapeutics and vaccines. Recently approved COVID-19 mRNA vaccines also utilize
the cellular machinery, to produce antigens and trigger immune responses. The design features of
the mRNA vaccines are critical to efficient mRNA translation in ribosomes, and are directly related
to the vaccine’s efficacy, safety, and immunogenicity. This review describes recent knowledge of
how the SARS-CoV-2 virus’ genomic characteristics interfere with ribosomal function and mRNA
translation. In addition, we discuss the current learning of the design features of mRNA vaccines and
their impacts on translational activity in ribosomes. The understanding of ribosomal interactions
with the virus and mRNA vaccines offers the foundation for antiviral therapeutic discovery and
continuous mRNA vaccine optimization to lower the dose, to increase durability and/or to reduce
adverse effects.
Keywords: ribosome; SARS-CoV-2; COVID-19 mRNA vaccines
1. Introduction
SARS-CoV-2 is the causing pathogen of the COVID-19 pandemic that has resulted in
more than 250 million cases and 5 million deaths [
1
–
3
]. Viruses employ the host cellular
translation machinery to synthesize their own proteins. Consequently, they have devel-
oped specialized mechanisms to commandeer the host machinery. SARS-CoV-2 uses a
multipronged procedure to manipulate host cellular machinery, to reduce global protein
translation and engage cellular resources in order to regulate their own protein production.
As the factory for protein synthesis in human cells, ribosomes play a critical role in infection
and human antiviral responses.
Each human ribosome consists of 2 unequal sized subunits; one is a 40S small subunit
and the other one is a large subunit (60S) [
4
]. The 40S small subunit is the decoding site that
consists of the 18S ribosomal RNA (rRNA) and 33 proteins. At the decoding site, sequence
information of the messenger RNA (mRNA) is translated into a protein sequence. The 60S
subunit is the peptidyl transferase center that harbors 28S, 5.8S, and 5S rRNAs, along with
47 proteins [
4
]. The peptidyl transferase catalyzes the peptide bond formation between
amino acids of the nascent protein at the 60S subunit [
5
]. The protein synthesis process
begins with translation initiation, a highly ordered process that regulates mRNA translation,
which is followed by elongation, during which a newly translated amino acid is added to
Life 2022, 12, 57. https://doi.org/10.3390/life12010057 https://www.mdpi.com/journal/life
Life 2022, 12, 57 2 of 14
the growing protein chain. The process is terminated when the ribosome completes the
translation with a stop codon. The protein synthesis process is an energy-intensive and
tightly regulated cellular process, with ribosome recycling being the last major step of this
process [6].
2. SARS-CoV-2 Interfere with Ribosome mRNA Translation
The SARS-CoV-2 is a novel beta-coronavirus, a category that also includes two already
known virulent coronaviruses, namely SARS-CoV-1 and MERS-CoV, that have resulted
in serious outbreaks in 2002 and 2012, respectively [
7
]. SARS-CoV-2 is an enveloped,
positive-sense and single-stranded RNA virus; its genome is approximately 30 kb in length.
The SARS-CoV-2 genome comprises a 5
0
-cap, 5
0
untranslated region (5
0
UTR), followed
by -replicase (ORF1a/ORF1b)-Spike (S)-Envelope (E)-Membrane (M)-Nucleocapsid (N)-3
0
UTR-poly(A) tail [
8
] (Figure 1). Though it shares more than 80% homology with SARS-CoV-
1 and ~50% with MERS-CoV, the mortality rates of these infections are slightly different,
ranging from 15% for SARS-CoV-1 and 34.4–37% for MERS-CoV, to around 2–13% for
SARS-CoV-2 [1,9–12].
Life 2022, 11, x FOR PEER REVIEW 2 of 15
translation, which is followed by elongation, during which a newly translated amino acid
is added to the growing protein chain. The process is terminated when the ribosome com-
pletes the translation with a stop codon. The protein synthesis process is an energy-inten-
sive and tightly regulated cellular process, with ribosome recycling being the last major
step of this process [6].
2. SARS-CoV-2 Interfere with Ribosome mRNA Translation
The SARS-CoV-2 is a novel beta-coronavirus, a category that also includes two al-
ready known virulent coronaviruses, namely SARS-CoV-1 and MERS-CoV, that have re-
sulted in serious outbreaks in 2002 and 2012, respectively [7]. SARS-CoV-2 is an envel-
oped, positive-sense and single-stranded RNA virus; its genome is approximately 30 kb
in length. The SARS-CoV-2 genome comprises a 5′-cap, 5′ untranslated region (5′ UTR),
followed by -replicase (ORF1a/ORF1b)-Spike (S)-Envelope (E)-Membrane (M)-Nucle-
ocapsid (N)-3′ UTR-poly(A) tail [8] (Figure 1). Though it shares more than 80% homology
with SARS-CoV-1 and ~50% with MERS-CoV, the mortality rates of these infections are
slightly different, ranging from 15% for SARS-CoV-1 and 34.4–37% for MERS-CoV, to
around 2–13% for SARS-CoV-2 [1,9–12].
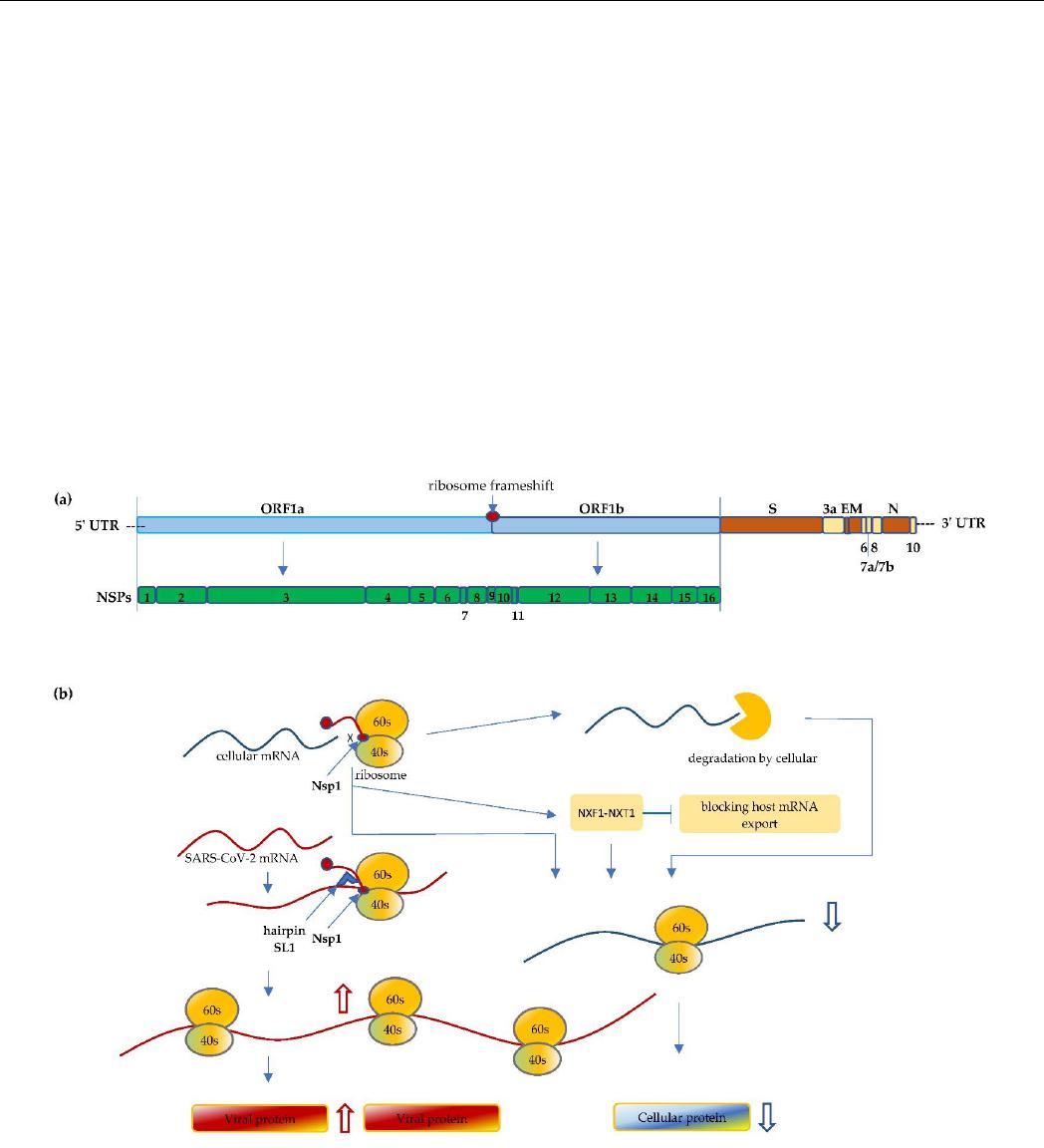
Figure 1. (a) Schematic illustration of the SARS-CoV-2 genome. The size of the coronavirus genome
is approximately 30 kb in length and comprises a 5′-cap, 5′ untranslated region (5′ UTR),
ORF1a/ORF1b, spike (S), envelope (E), membrane (M), nucleocapsid (N), and 3′ UTR-poly(A) tail.
The first ORF comprises of an approximated 2/3 of the genome that encodes the nonstructural pro-
teins (Nsp1 to Nsp16). (b) Nsp1 of SARS-CoV-2 binds to the 40S mRNA and block the mRNA en-
trance channel. Following viral infection, SARS-CoV-2 performs a multifaceted hijack on host ma-
chinery, including blocking the mRNA entry channel, accelerating host mRNA degradation, and
Figure 1.
(
a
) Schematic illustration of the SARS-CoV-2 genome. The size of the coronavirus genome is
approximately 30 kb in length and comprises a 5
0
-cap, 5
0
untranslated region (5
0
UTR), ORF1a/ORF1b,
spike (S), envelope (E), membrane (M), nucleocapsid (N), and 3
0
UTR-poly(A) tail. The first ORF
comprises of an approximated 2/3 of the genome that encodes the nonstructural proteins (Nsp1 to
Nsp16). (
b
) Nsp1 of SARS-CoV-2 binds to the 40S mRNA and block the mRNA entrance channel.
Following viral infection, SARS-CoV-2 performs a multifaceted hijack on host machinery, including
blocking the mRNA entry channel, accelerating host mRNA degradation, and inhibiting host mRNA
nucleus export. Furthermore, Nsp1 interacts with 5
0
UTR of SARS-CoV-2 and facilitates the translation
of its own protein, resulting in viral replication and protein accumulation, and inhibiting anti-viral
immune responses.

Life 2022, 12, 57 3 of 14
At the 5
0
end of SARS-CoV-2 genome, it starts with two large overlapping ORFs (ORF1a
and ORF1b) representing two-thirds of its genome, with the other one-third encoding the
structural proteins and accessory proteins [
13
,
14
]. Upon entering host cells, the ORF1a and
1b of the viral genomic RNA are translated and continually produce polyprotein, which is
then cleaved into functional NSPs. These NSPs play pivotal roles, like evading the host
immune system. Multiple NSPs interact with each other or form complex structures to
modify cellular conditions for efficient mRNA translation and viral replication [
15
]. The
440–500 kDa Polyprotein 1a (pp1a) is first translated from ORF1a and then cleaved from
Nsp1 to Nsp11. Between ORF1a and 1b, the programmed ribosomal -1 frameshift (PRF)
of the reading frame immediately takes place preceding the stop codon of ORF1a, which
enables the downstream translation of pp1ab from ORF1b. The pp1ab is subsequently
processed into functional Nsp1 through Nsp16 [16].
The PRF, a well-preserved process among coronaviruses, is essential for functioning
translation of NSPs. An efficient PRF engages a conserved slippery sequence (U_UUA_AAC)
that changes the reading frame to UUU_AAA_C after the frameshifting [
17
]. When a ri-
bosome approaches the slippery site during translation, a stimulatory RNA folds into
a stable pseudoknot structure that slows down translation and promotes the PRF [
17
].
Importantly, Nsp12 and downstream NSPs that are involved in RNA capping, modification,
and proofreading, rely on the PRF process as they are translated after the frameshifting [
17
].
It has been reported that multiple factors, including, for example, the position of the ORF1a
stop codon, and interactions between ribosomal tunnels and RNA elements, including
pseudoknot and nascent chain, modulate the optimum efficiency of frameshifting [
17
]. In
addition to RNA regulation, the zinc-finger antiviral protein (ZAP-s) has been observed
to interact with viral RNA and interfere with PRF [
18
]. PRF is one of the crucial steps in
ribosome translation of virus genomes and viral replication, and thus presents a viable
potential target for antiviral intervention therapeutics [17,19,20].
Among the 16 NSPs, Nsp1 is one of the first functional coronaviral nonstructural
proteins translated in infected cells. Nsp1 is a protein comprised of 180 amino acids that
targets cellular processes to inhibit translation, triggers host mRNA cleavage and decay,
and down-regulates type I interferon (IFN) response [
21
–
24
]. NSP1 is a major virulence
factor that is essential for viral replication, and is thus emphasized here [15,19,22,25].
Known as the host shutoff factor, Nsp1 efficiently interacts with an array of different
ribosomal states, resulting in a shutdown of host protein production [
15
,
22
]. The 40S
subunit plays a critical function in the highly regulated translation initiation process, in
which it binds initiation factors to form 43S pre-initiation complexes and facilitates scanning
of the 5
0
UTR to the AUG start codon [
26
]. It has been reported that Nsp1 manipulates the
ribosome at the translation initiation step and stalls canonical mRNA translation directly
through binds to the ribosomes’ 40S subunit, the 43S pre-initiation complex, and the 80S
non-translating ribosome [
15
,
21
,
22
]. The C-terminal domain of the Nsp1 protein folds
into two helices that physically block mRNA accommodation at the entrance channel and
shut down host mRNA translation [
15
]. Mutants of the C-terminal domain led to the
abolishment of the Nsp1’s ability to bind 40S subunits, indicating that the helices area,
specifically aa 154–165 and 171–179, are crucial for Nsp1–ribosome interactions [15,25].
In addition to inhibition of the host mRNA translation, SARS-CoV-2 has developed
manifold instruments, including, for example, the degradation of host mRNA and the
blocking of host mRNA export. Nsp1 facilitates accelerated host mRNA degradation,
through mRNA endonucleolytic cleavage in the 5
0
-UTR of the host mRNA. In particular,
the R124 and K125 aa sites of the Nsp1 protein play a pivotal role in this cleavage [
25
]. The
accelerated degradation of cytosolic cellular mRNAs is a significant part of remodeling the
mRNA pool, facilitating the viral takeover of the mRNA pool in infected cells [
14
,
27
,
28
].
Furthermore, Nsp1 binds to the host mRNA export factor to interact with the NXF1-NXT1
receptor resulting in the inhibition of mRNA nuclear export. As a result, host mRNA is
retained in the nucleus [24] (Figure 1).

Life 2022, 12, 57 4 of 14
While shutting off host mRNA translation, SARS-CoV-2 orchestrates its own viral
translation without inhibition under the ribosome blockade condition, which is achieved
through the highly ordered process by multifunctional NSPs. Despite the Nsp1 bound on
the ribosome, Nsp1 of SARS-CoV-2 can recognize and accommodate its own mRNA and
proceed to initiate the translation. The 5
0
UTR of SARS-CoV-2 forms a unique cis-acting
RNA hairpin SL1 structure that plays a prominent role in this evasion. The SL1 structure
interacts with Nsp1, which frees the mRNA accommodation channel and promotes viral
translation [
28
]. It has been reported that the binding affinity between Nsp1 and SL1 is
relatively high, which facilitates the recruitment of the 40S ribosomal subunit and favors
the viral translation [
29
]. In addition, the viral mRNA is protected from endonucleolytic
mRNA cleavage and degradation. The interaction of Nsp1 and 5
0
UTR of viral RNA can
also contribute the resistant to the cleavage [
29
]. Another mechanism is the interaction
between Nsp10, Nsp14, and Nsp16, which forms a complex that facilitates SARS-CoV-
2 mRNA capping and proof-reading, providing a significant contribution in protecting
SARS-CoV-2 mRNA from accelerated degradation [
13
]. Through a collective mechanism,
Nsp1 of SARS-CoV-2 is vital for viral replication as it not only hampers the translation of
cellular transcripts, shutting off host protein production, but also has the ability to recruit
the ribosome in order to efficiently translate the viral mRNA to allow the expression of
viral genes [15].
Type I interferon induction and innate interferon response represent one of the major
innate antiviral host defenses against viral infections [
30
]. SARS-CoV-2 efficiently sup-
presses the IFN-I signaling, likely mediated by the inhibition of STAT1 and STAT2, resulting
in lack of efficient IFN-dependent antiviral innate immune responses. In addition to the
shutting-off of host mRNA translation, the inhibition of INF-I antiviral responses leads to
higher viral replication, viral protein accumulation, and pathogenesis [
30
]. Collectively, a
fully functional Nsp1 is necessary for virulence; thus, the targeting of Nsp1 proteins and
the Nsp1–ribosome interactions presents an attractive therapeutic opportunity for future
studies [13,22,28].
3. Mutations Impact Replication and Virulence
Although SARS-CoV-2 has proof-reading processes, mutations arise naturally during
viral replication, which causes new variants to form. Since late 2020, several novel variants
have been named SARS-CoV-2 variants of concern/interest (VOC/I) due to their greater
risk of enhanced transmissibility, pathogenicity and/or ability to evade host response [
31
].
The Alpha (B.1.1.7) variant, first detected in England in September 2020, appears to have
a higher reproduction number and transmits more efficiently from person to person [
32
].
The Beta (B.1.351) variants, first reported in South America, and Gamma (P.1) variants
share some of the same genetic changes that are associated with the increased transmission,
and higher viral load. It is reported that these mutations lead to immune escape from
neutralizing antibodies [
33
,
34
]. The Delta (B.1.617.2) variant is believed to be highly
transmissible with more than twice as the original strain of SARS-CoV-2 [
35
]. Since first
appearing in India in late 2020, it has spread worldwide and became the dominant variant
of SARS-CoV-2 virus in the U.S. in late 2021 [
36
]. The recent emergence of Omicron has
raised significant concern due to its extensive mutations, including more than 30 mutations
on the Spike protein [37].
The virus is able to mutate in a dangerous and clever way to become structurally more
infectious or cause more severe disease, in which multiple mechanisms may be operating.
The major VOCs have shared mutations in the spike protein of SARS-CoV-2 genome,
mostly on the S1 subunit, which is the unit that possesses the receptor-binding domain
(RBD) that binds to cellular receptor ACE2 through six key amino acid residues [
38
]
(Figure 2). N501 on the RBD domain is one of these amino acids that has a specific
interaction with ACE2 receptor. Observed in three variants (Alpha, Beta, and Gamma), the
N501Y mutation raises considerable concern due to its greater ACE2 binding affinity and
enhanced transmissibility [
39
,
40
]. The Beta variant carries K417N and E484K mutations
Life 2022, 12, 57 5 of 14
while Gamma shares E484K, along with the mutation of K417 to K417T [
40
]. These S1
mutations increase the binding affinity to the ACE2 receptor, thereby possibly enhancing
transmissibility, which can affect disease severity and clinical outcomes [38].
The Delta variant has picked up new mutations, including multiple mutations (L452R,
E484Q, T478K) in the S1 subunit, resulting in a viral load which is more than 1000 times
higher than the original strain infections [
35
]. The Delta variant is broken into a few sub-
types that are being classified as Delta Plus, which has drawn greater attention recently [
41
].
In addition to the Delta mutations, the Delta-AY.1 variant carries K417N mutation that
can interact with N501Y, which can increase the binding affinity with ACE2 receptor and
possibly reduce neutralizing antibody susceptibility [
41
]. D614G, a non-RBD site mutation
on spike proteins, represented in more than 90% of prevalent variants, has become the
most predominant mutation since it first emerged in early 2020 [
38
]. It is reported that the
D614G is related to increase spike density and cell entry, but does not directly increase the
ACE2 receptor binding affinity or reduce the susceptibility of neutralizing antibodies for
the virion [42].
Life 2022, 11, x FOR PEER REVIEW 5 of 15
interaction with ACE2 receptor. Observed in three variants (Alpha, Beta, and Gamma),
the N501Y mutation raises considerable concern due to its greater ACE2 binding affinity
and enhanced transmissibility [39,40]. The Beta variant carries K417N and E484K muta-
tions while Gamma shares E484K, along with the mutation of K417 to K417T [40]. These
S1 mutations increase the binding affinity to the ACE2 receptor, thereby possibly enhanc-
ing transmissibility, which can affect disease severity and clinical outcomes [38].
The Delta variant has picked up new mutations, including multiple mutations
(L452R, E484Q, T478K) in the S1 subunit, resulting in a viral load which is more than 1000
times higher than the original strain infections [35]. The Delta variant is broken into a few
subtypes that are being classified as Delta Plus, which has drawn greater attention re-
cently [41]. In addition to the Delta mutations, the Delta-AY.1 variant carries K417N mu-
tation that can interact with N501Y, which can increase the binding affinity with ACE2
receptor and possibly reduce neutralizing antibody susceptibility [41]. D614G, a non-RBD
site mutation on spike proteins, represented in more than 90% of prevalent variants, has
become the most predominant mutation since it first emerged in early 2020 [38]. It is re-
ported that the D614G is related to increase spike density and cell entry, but does not
directly increase the ACE2 receptor binding affinity or reduce the susceptibility of neu-
tralizing antibodies for the virion [42].
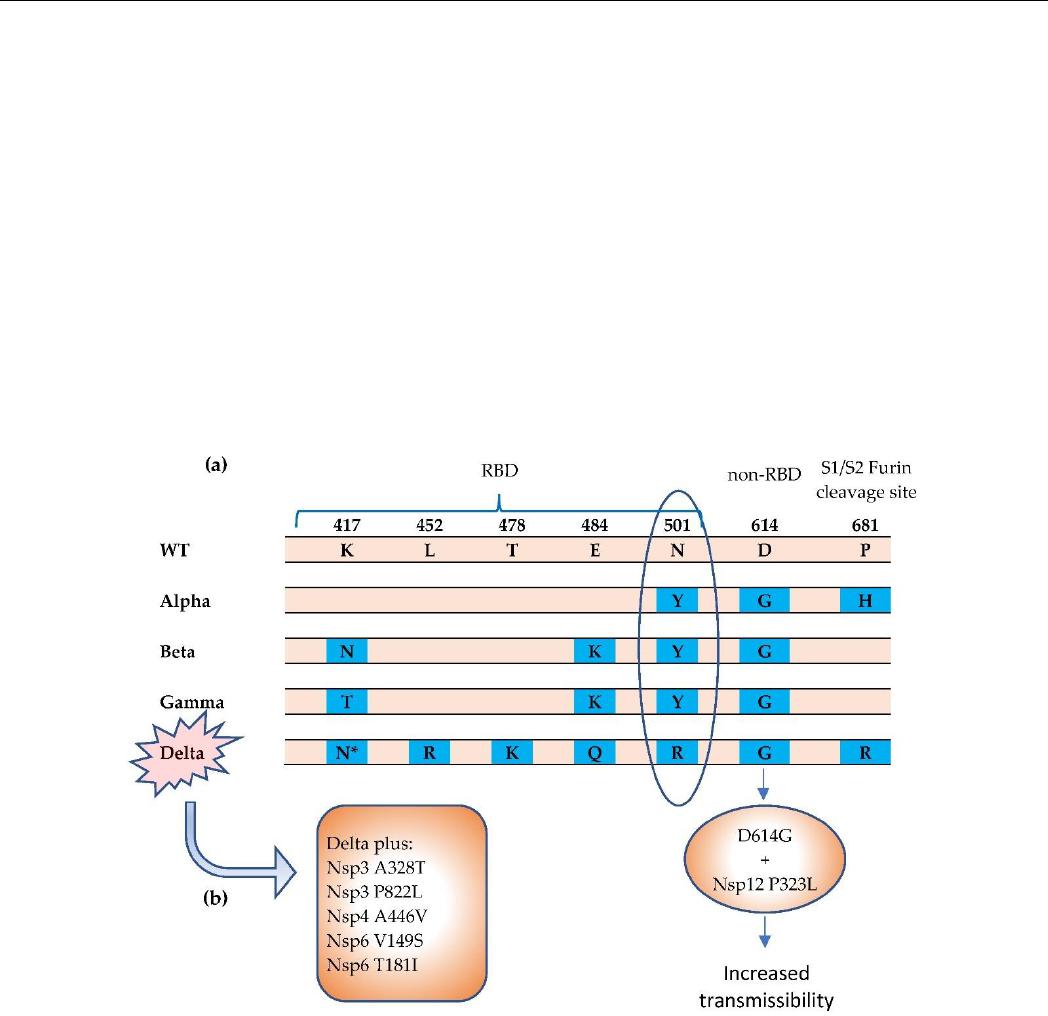
Figure 2. The Alpha, Beta, Gamma, and Delta variants have been termed SARS-CoV-2 COVs. (a)
Notable mutations in spike protein [38–40]. The N501Y mutation results in greater affinity for ACE2
receptor, which can increase transmissibility. (b) The Delta variant carries a more diverse repertoire
of mutations [43,44]. The Delta Plus variant carries increased mutations in NSPs. * = K417N. The
K417N mutation is significantly more prevalent in the Delta Plus (AY.1 or B.1.617.2.1) variant than
in the Delta (B.1.617.2) variant. RBD = receptor binding domain.
Although our understanding of the functional consequences of spike mutations is
rapidly expanding, the mutations in NSPs, 3′ structural proteins, and accessory proteins
are relatively under-investigated despite playing a significant role in virus translation,
replication, and host immune suppression.
Figure 2.
The Alpha, Beta, Gamma, and Delta variants have been termed SARS-CoV-2 COVs.
(
a
) Notable mutations in spike protein [
38
–
40
]. The N501Y mutation results in greater affinity for
ACE2 receptor, which can increase transmissibility. (
b
) The Delta variant carries a more diverse
repertoire of mutations [
43
,
44
]. The Delta Plus variant carries increased mutations in NSPs.
* = K417N
.
The K417N mutation is significantly more prevalent in the Delta Plus (AY.1 or B.1.617.2.1) variant
than in the Delta (B.1.617.2) variant. RBD = receptor binding domain.
Although our understanding of the functional consequences of spike mutations is
rapidly expanding, the mutations in NSPs, 3
0
structural proteins, and accessory proteins
are relatively under-investigated despite playing a significant role in virus translation,
replication, and host immune suppression.
NSP1 is a highly conserved protein with few mutations that have been identified [
25
].
The two helices structure, including in particular the KH motif K164 and H165 in C-terminal
of the SARS-CoV-2 Nsp1, is crucial for ribosome binding and the inhibition of translation
initiation. [
15
]. However, it is interesting that a deletion hotspot at the 500–532 locus of
the Nsp1 N-terminus coding region has been identified, which has been detected in 37
